What is BSL?
Bayesian Synthetic Likelihood is a "likelihood-free" inference method — like ABC-SMC
(notebook 02), it does not require a mathematical formula for the probability of the data
given the parameters. Instead, it uses simulation to approximate that probability.
The biological analogy: imagine you want to know whether a particular
combination of mutation rate, DFE shape, and other parameters could have produced the
fitness trajectory you observed in a population. You cannot solve this analytically —
the mutation-selection process is too complex. So instead, you run 50 simulations at those
parameter values and look at the results. If the observed data looks "typical" compared to
those 50 simulations, the parameters are plausible. If the observed data is an extreme
outlier, those parameters are unlikely.
BSL formalizes this intuition: it fits a multivariate normal (bell curve) distribution to the
summary statistics from the 50 simulations, then evaluates how probable the observed
statistics are under that bell curve. This gives a number — the synthetic
likelihood — that can be plugged into standard MCMC machinery to explore parameter
space and build a posterior distribution.
How does BSL differ from ABC-SMC?
| Feature |
ABC-SMC (notebook 02) |
BSL (this notebook) |
| Core question |
"Are these simulations close enough to the data?" |
"What is the probability of the data under a normal model fitted to these simulations?" |
| Epsilon threshold |
Required — defines "close enough" (arbitrary choice) |
Not needed — uses a proper likelihood instead |
| Convergence diagnostics |
Heuristic (particle diversity, ESS of weights) |
Formal MCMC diagnostics (R-hat, ESS, trace plots) |
| Sampling mechanism |
Sequential importance sampling with particles |
MCMC random walk (BlackJAX adaptive Metropolis) |
| Key assumption |
Good distance metric between simulations and data |
Summary statistics are approximately multivariate normal |
| Computational cost per step |
One simulation per particle |
50 simulations per MCMC step (more expensive, but fewer steps needed) |
Why use both? If ABC-SMC and BSL agree on the inferred parameter values,
we have strong evidence that the results are robust — they are not artifacts of either
method's assumptions. This is especially important when the conclusions have biological
significance, such as determining whether a population sits above or below the mutational
meltdown threshold in the Basener & Sanford framework.
MCMC diagnostics: why they matter
A key advantage of BSL over ABC-SMC is that it produces standard MCMC chains with formal
convergence diagnostics. These tell us whether we can trust the posterior:
- R-hat (Gelman-Rubin statistic): Compares variance within each chain to
variance between the two chains. Values near 1.0 (below 1.05) mean both
chains have converged to the same distribution — they agree on the answer. In plain
language: the sampler started from two different places in parameter space and arrived at
the same conclusions. Values above 1.1 indicate the chains have not yet converged and the
results are unreliable.
- ESS (Effective Sample Size): Because MCMC samples are correlated (each
step depends on the previous one), 500 raw samples may contain only ~100–200
effectively independent draws. ESS above 100 is generally adequate for estimating posterior
means and credible intervals. Low ESS suggests the chain needs to run longer or the
proposal needs better tuning.
- Acceptance rate: The fraction of proposed MCMC moves that were accepted.
Rates of 20–40% are typical for well-tuned random-walk Metropolis in multiple
dimensions. Very low rates (<10%) mean the proposals are too large; very high rates
(>90%) mean the proposals are too small and the chain is barely moving.
In plain language: convergence means the sampler has explored enough of the
five-dimensional parameter space to give reliable answers. It has visited all the plausible
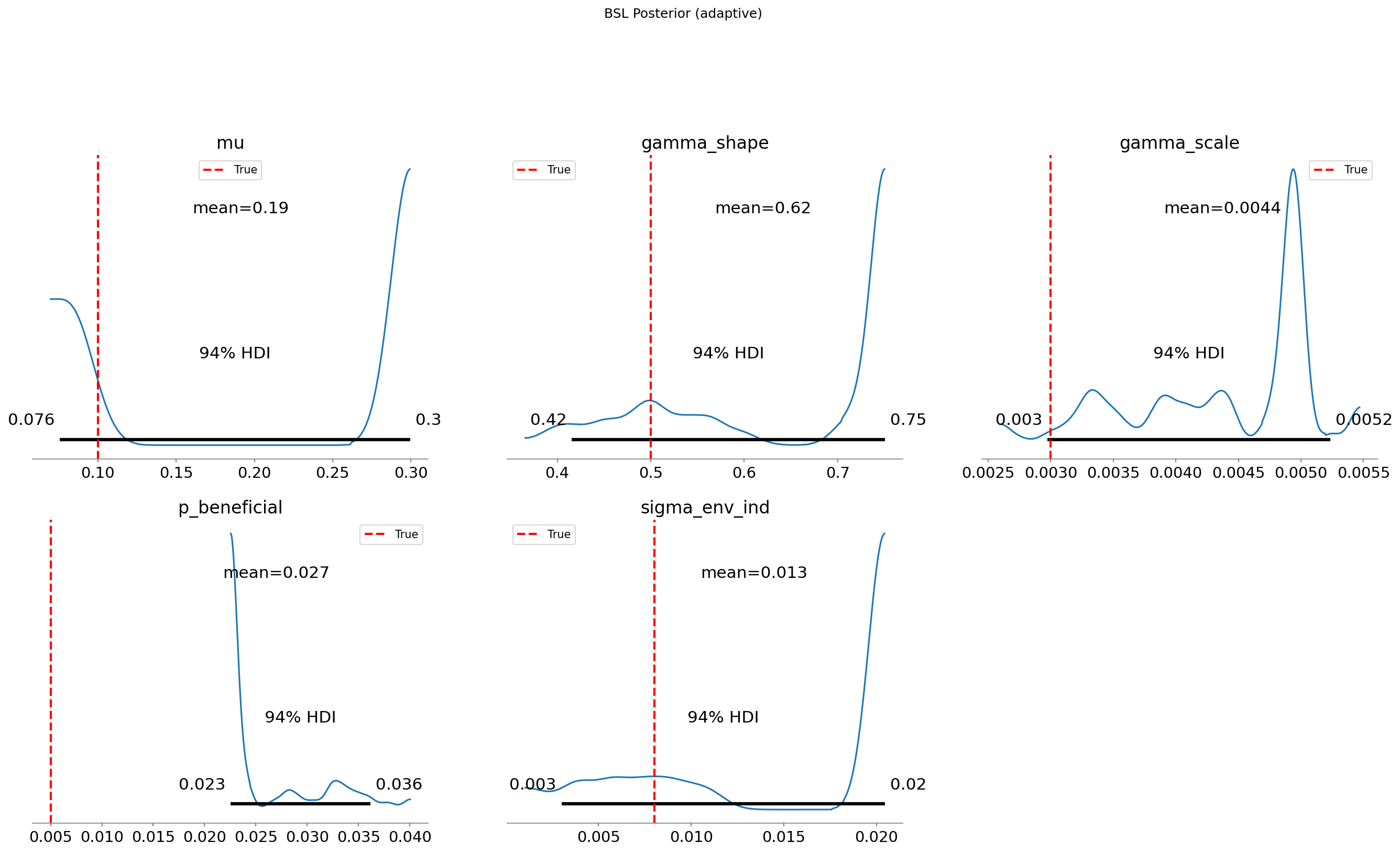
parameter combinations, not just a small corner. Good diagnostics (R-hat near 1.0,
ESS > 100) are our assurance that running the sampler longer would not substantially change
the posterior distributions shown above.